artan.filter package¶
Submodules¶
artan.filter.filter_params module¶
-
class
artan.filter.filter_params.HasCalculateLoglikelihood[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param for enabling loglikelihood calculation.
-
calculateLoglikelihood= Param(parent='undefined', name='calculateLoglikelihood', doc='When true, loglikelihood of residual will be calculated & added to output DataFrame. Default is false')¶
-
-
class
artan.filter.filter_params.HasCalculateMahalanobis[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param for enabling mahalanobis calculation.
-
calculateMahalanobis= Param(parent='undefined', name='calculateMahalanobis', doc='When true, mahalanobis distance of residual will be calculated & added to output DataFrame.Default is false.')¶
-
-
class
artan.filter.filter_params.HasCalculateSlidingLikelihood[source]¶ Bases:
pyspark.ml.param.ParamsMixin param for enabling sliding likelihood calculation
-
calculateSlidingLikelihood= Param(parent='undefined', name='calculateSlidingLikelihood', doc='When true, sliding likelihood sum of residual will be calculated & added to output DataFrame. Default is false')¶
-
-
class
artan.filter.filter_params.HasControlCol[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param for control column.
-
controlCol= Param(parent='undefined', name='controlCol', doc='Column name for specifying control vector')¶
-
-
class
artan.filter.filter_params.HasControlFunctionCol[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param for control function column.
-
controlFunctionCol= Param(parent='undefined', name='controlFunctionCol', doc='Column name for specifying control matrix')¶
-
-
class
artan.filter.filter_params.HasFadingFactor[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param fading factor.
-
fadingFactor= Param(parent='undefined', name='fadingFactor', doc='Factor controlling the weight of older measurements. With larger factor, more weights willbe given to recent measurements. Typically, should be really close to 1')¶
-
-
class
artan.filter.filter_params.HasInitialStateCovariance[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param initial covariance matrix.
-
getInitialStateCovariance()[source]¶ Gets the value of initial covariance matrix or its default value.
-
initialStateCovariance= Param(parent='undefined', name='initialStateCovariance', doc='Initial covariance matrix')¶
-
-
class
artan.filter.filter_params.HasInitialStateCovarianceCol[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param for initial covariance column.
-
getInitialStateCovarianceCol()[source]¶ Gets the value of initial covariance column or its default value.
-
initialStateCovarianceCol= Param(parent='undefined', name='initialStateCovarianceCol', doc='Column name for initial state covariance matrix.')¶
-
-
class
artan.filter.filter_params.HasInitialStateDistributionCol[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param for initial state distribution column.
-
getInitialStateDistributionCol()[source]¶ Gets the value of initial distribution column or its default value.
-
initialStateDistributionCol= Param(parent='undefined', name='initialStateDistributionCol', doc='Parameter for initial state distribution as struct col')¶
-
-
class
artan.filter.filter_params.HasInitialStateMean[source]¶ Bases:
pyspark.ml.param.ParamsMixin for initial state vector.
-
initialStateMean= Param(parent='undefined', name='initialStateMean', doc='Initial state vector')¶
-
-
class
artan.filter.filter_params.HasInitialStateMeanCol[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param for initial state column.
-
initialStateMeanCol= Param(parent='undefined', name='initialStateMeanCol', doc='Column name for initial state vector.')¶
-
-
class
artan.filter.filter_params.HasMeasurementCol[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param for measurement column.
-
measurementCol= Param(parent='undefined', name='measurementCol', doc='Column name for measurement vector. Missing measurements are allowed with nulls in the data')¶
-
-
class
artan.filter.filter_params.HasMeasurementModel[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param measurement model matrix.
-
measurementModel= Param(parent='undefined', name='measurementModel', doc='Measurement matrix, when multiplied with the state it should give the measurement vector')¶
-
-
class
artan.filter.filter_params.HasMeasurementModelCol[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param for measurement model column.
-
measurementModelCol= Param(parent='undefined', name='measurementModelCol', doc='Column name for specifying measurement model from input DataFrame rather thana constant measurement model for all filters')¶
-
-
class
artan.filter.filter_params.HasMeasurementNoise[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param measurement noise matrix.
-
measurementNoise= Param(parent='undefined', name='measurementNoise', doc='Measurement noise matrix')¶
-
-
class
artan.filter.filter_params.HasMeasurementNoiseCol[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param for measurement noise column.
-
measurementNoiseCol= Param(parent='undefined', name='measurementNoiseCol', doc='Column name for specifying measurement noise from input DataFrame rather thana constant measurement noise for all filters')¶
-
-
class
artan.filter.filter_params.HasMeasurementSize[source]¶ Bases:
pyspark.ml.param.ParamsMixin for measurement vector size
-
measurementSize= Param(parent='undefined', name='measurementSize', doc='Size of the measurement vector')¶
-
-
class
artan.filter.filter_params.HasMultiStepPredict[source]¶ Bases:
pyspark.ml.param.ParamsMixin for multi step predict
-
multiStepPredict= Param(parent='undefined', name='multiStepPredict', doc='Number of predict steps after a predict&update cycle')¶
-
-
class
artan.filter.filter_params.HasMultipleModelAdaptiveEstimationEnabled[source]¶ Bases:
pyspark.ml.param.Params-
multipleModelAdaptiveEstimationEnabled= Param(parent='undefined', name='multipleModelAdaptiveEstimationEnabled', doc='Flag for enabling Multiple Model Adaptive Estimation (MMAE) output mode. When enabled,MMAE mode outputs a single state estimate from the output of all kalman states of the transformer.States are weighted based on their sliding likelihood')¶
-
-
class
artan.filter.filter_params.HasMultipleModelMeasurementWindowDuration[source]¶ Bases:
pyspark.ml.param.Params-
getMultipleModelMeasurementWindowDuration()[source]¶ Gets the value of mmae measureent window duration
-
multipleModelMeasurementWindowDuration= Param(parent='undefined', name='multipleModelMeasurementWindowDuration', doc='Window duration for grouping measurements in same window for MMAE filter aggregation')¶
-
-
class
artan.filter.filter_params.HasOutputSystemMatrices[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param for enabling the output of system matrices along with the state.
-
outputSystemMatrices= Param(parent='undefined', name='outputSystemMatrices', doc='When true, the system matrices will be added to output DataFrame. Default is false')¶
-
-
class
artan.filter.filter_params.HasProcessModel[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param process model matrix.
-
processModel= Param(parent='undefined', name='processModel', doc='Process model matrix, transitions the state to the next state when applied')¶
-
-
class
artan.filter.filter_params.HasProcessModelCol[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param for process model column.
-
processModelCol= Param(parent='undefined', name='processModelCol', doc='Column name for specifying process model from input DataFrame rather thana constant measurement model for all filters')¶
-
-
class
artan.filter.filter_params.HasProcessNoise[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param process noise matrix.
-
processNoise= Param(parent='undefined', name='processNoise', doc='Process noise matrix')¶
-
-
class
artan.filter.filter_params.HasProcessNoiseCol[source]¶ Bases:
pyspark.ml.param.ParamsMixin for param for process noise column.
-
processNoiseCol= Param(parent='undefined', name='processNoiseCol', doc='Column name for specifying process noise from input DataFrame rather thana constant measurement noise for all filters')¶
-
-
class
artan.filter.filter_params.HasSlidingLikelihoodWindow[source]¶ Bases:
pyspark.ml.param.ParamsMixin param for sliding likelihood window duration
-
slidingLikelihoodWindow= Param(parent='undefined', name='slidingLikelihoodWindow', doc='Number of consecutive measurements to include in the total likelihood calculation')¶
-
-
class
artan.filter.filter_params.HasStateSize[source]¶ Bases:
pyspark.ml.param.ParamsMixin for state vector size
-
stateSize= Param(parent='undefined', name='stateSize', doc='Size of the state vector')¶
-
-
class
artan.filter.filter_params.KalmanFilterParams[source]¶ Bases:
artan.filter.filter_params.HasInitialStateMean,artan.filter.filter_params.HasInitialStateCovariance,artan.filter.filter_params.HasInitialStateMeanCol,artan.filter.filter_params.HasInitialStateCovarianceCol,artan.filter.filter_params.HasInitialStateDistributionCol,artan.filter.filter_params.HasProcessModel,artan.filter.filter_params.HasFadingFactor,artan.filter.filter_params.HasMeasurementModel,artan.filter.filter_params.HasMeasurementNoise,artan.filter.filter_params.HasProcessNoise,artan.filter.filter_params.HasMeasurementCol,artan.filter.filter_params.HasMeasurementModelCol,artan.filter.filter_params.HasMeasurementNoiseCol,artan.filter.filter_params.HasProcessModelCol,artan.filter.filter_params.HasProcessNoiseCol,artan.filter.filter_params.HasControlCol,artan.filter.filter_params.HasControlFunctionCol,artan.filter.filter_params.HasCalculateMahalanobis,artan.filter.filter_params.HasCalculateLoglikelihood,artan.filter.filter_params.HasOutputSystemMatrices,artan.filter.filter_params.HasCalculateSlidingLikelihood,artan.filter.filter_params.HasSlidingLikelihoodWindow,artan.filter.filter_params.HasMultiStepPredict,artan.filter.filter_params.HasStateSize,artan.filter.filter_params.HasMeasurementSizeMixin for kalman filter parameters
-
setCalculateLogLikelihood()[source]¶ Optionally calculate loglikelihood of each measurement & add it to output dataframe. Loglikelihood is calculated from residual vector & residual covariance matrix.
Not enabled by default.
Returns: KalmanFilter
-
setCalculateMahalanobis()[source]¶ Optionally calculate mahalanobis distance metric of each measurement & add it to output dataframe. Mahalanobis distance is calculated from residual vector & residual covariance matrix.
Not enabled by default.
Returns: KalmanFilter
-
setCalculateSlidingLikelihood()[source]¶ Optionally calculate a sliding likelihood across consecutive measurements
Default is false
Returns: KalmanFilter
-
setControlCol(value)[source]¶ Set the column for input control vectors.
Control vectors should have compatible size with control function (controlVectorSize). The product of control matrix & vector should produce a vector with stateSize. null values are allowed, which will result in state transition without control input.
Parameters: value – String Returns: KalmanFilter
-
setControlFunctionCol(value)[source]¶ Set the column for input control matrices.
Control matrices should have dimensions (stateSize, controlVectorSize). null values are allowed, which will result in state transition without control input
Parameters: value – String Returns: KalmanFilter
-
setFadingFactor(value)[source]¶ Fading factor for giving more weights to more recent measurements. If needed, it should be greater than one. Typically set around 1.01 ~ 1.05. Default is 1.0, which will result in equally weighted measurements.
Parameters: value – Float >= 1.0 Returns: KalmanFilter
-
setInitialStateCovariance(value)[source]¶ Set the initial covariance matrix with dimensions (stateSize, stateSize)
It will be applied to all states. If the state timeouts and starts receiving measurements after timeout, it will again start from this initial covariance vector. Default is identity matrix. :param value: pyspark.ml.linalg.Matrix with dimensions (stateSize, stateSize) :return: KalmanFilter
-
setInitialStateCovarianceCol(value)[source]¶ Set the column corresponding to initial covariance matrix. Overrides setInitialCovariance parameter.
Parameters: value – String Returns: KalmanFilter
-
setInitialStateDistributionCol(value)[source]¶ Set the column corresponding to initial state distribution struct. The struct should have ‘mean’ field with a VectorType and ‘covariance’ field with a MatrixType. Overrides setInitialStateMeanCol and setInitialCovarianceCol parameters.
Parameters: value – String Returns: KalmanFilter
-
setInitialStateMean(value)[source]¶ Set the initial state vector with size (stateSize).
It will be applied to all states. If the state timeouts and starts receiving measurements after timeout, it will again start from this initial state vector. Default is zero.
Note that if this parameter is set through here, it will result in same initial state for all filters. For different initial states across filters, set the dataframe column for corresponding to initial state with setInitialStateCol.
Parameters: value – pyspark.ml.linalg.Vector with size (stateSize) Returns: KalmanFilter
-
setInitialStateMeanCol(value)[source]¶ Set the column corresponding to initial state vector. Overrides setInitialState setting.
Parameters: value – String Returns: KalmanFilter
-
setMeasurementCol(value)[source]¶ Set the column corresponding to measurements.
The vectors in the column should be of size (measurementSize). null values are allowed, which will result in only state prediction step.
Parameters: value – pyspark.ml.linalg.Vector with size measurementSize Returns: KalmanFilter
-
setMeasurementModel(value)[source]¶ Set default value for measurement model matrix with dimensions (stateSize, measurementSize) which maps states to measurement.
Note that if this parameter is set through here, it will result in same measurement model for all filters & measurements. For different measurement models across filters or measurements, set a dataframe column for measurement model from setMeasurementModelCol.
Default value maps the first state value to measurements.
Parameters: value – pyspark.ml.linalg.Matrix with dimensions (stateSize, measurementSize) Returns: KalmanFilter
-
setMeasurementModelCol(value)[source]¶ Set the column for input measurement model matrices
Measurement model matrices should have dimensions (stateSize, measurementSize)
Parameters: value – String Returns: KalmanFilter
-
setMeasurementNoise(value)[source]¶ Set default value for measurement noise matrix with dimensions (measurementSize, measurementSize).
Note that if this parameter is set through here, it will result in same measurement noise for all filters & measurements. For different measurement noise values across filters or measurements, set a dataframe column for measurement noise from setMeasurementNoiseCol.
Default is identity matrix.
Parameters: value – pyspark.ml.linalg.Matrix with dimensions (measurementSize, measurementSize) Returns: KalmanFilter
-
setMeasurementNoiseCol(value)[source]¶ Set the column for input measurement noise matrices.
Measurement noise matrices should have dimensions (measurementSize, measurementSize)
Parameters: value – String Returns: KalmanFilter
-
setMeasurementSize(value)[source]¶ Sets the measurement size vector. Only required apriori for MMAE mode.
Parameters: value – Integer Returns: KalmanFilter
-
setMultiStepPredict(value)[source]¶ Sets the param for running multiple prediction steps after measurement updates in Kalman filter. By default, a single prediction step is followed by a single measurement update step. If this parameter is set to a number n>0, then n-predict steps will be undertaken after measurement update steps.
Default is 0 :param value: Integer :return: KalmanFilter
-
setOutputSystemMatrices()[source]¶ Optionally add system matrices to output dataframe returned by the transformer.
Default is false
Returns: KalmanFilter
-
setProcessModel(value)[source]¶ Set default value for process model matrix with dimensions (stateSize, stateSize) which governs state transition.
Note that if this parameter is set through here, it will result in same process model for all filters & measurements. For different process models across filters or measurements, set a dataframe column for process model from setProcessModelCol.
Default is identity matrix.
Parameters: value – pyspark.ml.linalg.Matrix with dimensions (stateSize, stateSize) Returns: KalmanFilter
-
setProcessModelCol(value)[source]¶ Set the column for input process model matrices.
Process model matrices should have dimensions (stateSize, stateSize)
Parameters: value – String Returns: KalmanFilter
-
setProcessNoise(value)[source]¶ Set default value for process noise matrix with dimensions (stateSize, stateSize).
Note that if this parameter is set through here, it will result in same process noise for all filters & measurements. For different process noise values across filters or measurements, set a dataframe column for process noise from setProcessNoiseCol.
Default is identity matrix.
Parameters: value – pyspark.ml.linalg.Matrix with dimensions (stateSize, StateSize) Returns: KalmanFilter
-
setProcessNoiseCol(value)[source]¶ Set the column for input process noise matrices.
Process noise matrices should have dimensions (stateSize, stateSize)
Parameters: value – String Returns: KalmanFilter
-
artan.filter.least_mean_squares_filter module¶
-
class
artan.filter.least_mean_squares_filter.LeastMeanSquaresFilter[source]¶ Bases:
artan.state.stateful_transformer.StatefulTransformer,artan.filter.filter_params.HasInitialStateMean,artan.filter.filter_params.HasInitialStateMeanCol,artan.filter.least_mean_squares_filter._HasLearningRate,artan.filter.least_mean_squares_filter._HasRegularizationConstant,pyspark.ml.param.shared.HasLabelCol,pyspark.ml.param.shared.HasFeaturesCol,artan.utils.ArtanJavaMLReadable,pyspark.ml.util.JavaMLWritableNormalized Least Mean Squares filter, implemented with a stateful spark Transformer for running parallel filters /w spark dataframes. Transforms an input dataframe of observations to a dataframe of model parameters using stateful spark transformations, which can be used in both streaming and batch applications.
Belonging to stochastic gradient descent type of methods, LMS minimizes SSE on each measurement based on the expectation of steepest descending gradient.
Let
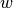 denote the model parameter vector,
denote the model parameter vector, 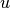 denote the features vector,
and
denote the features vector,
and 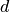 the label corresponding to
the label corresponding to 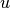 . Normalized LMS computes
. Normalized LMS computes 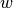 at step
at step 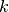 recursively by:
recursively by: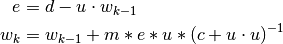
Where -
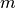 : Learning rate
-
: Learning rate
-  : Regularization constant
: Regularization constant-
setFeaturesCol(value)[source]¶ Set features column. Default is “features”
Parameters: value – String Returns: RecursiveLeastSquaresFilter
-
setInitialEstimate(value)[source]¶ Set initial estimate for model parameters. Default is zero vector.
Note that if this parameter is set through here, it will result in same initial estimate for all filters. For different initial estimates across filters, set the dataframe column for corresponding to initial estimate with setInitialEstimateCol.
Parameters: value – pyspark.ml.linalg.Vector with size (featuresSize) Returns: RecursiveLeastSquaresFilter
-
setInitialEstimateCol(value)[source]¶ Sets the column corresponding to initial estimates :param value: String :return: RecursiveLeastSquaresFilter
-
setLabelCol(value)[source]¶ Set label column. Default is “label”
Parameters: value – String Returns: RecursiveLeastSquaresFilter
-
artan.filter.linear_kalman_filter module¶
-
class
artan.filter.linear_kalman_filter.LinearKalmanFilter[source]¶ Bases:
artan.state.stateful_transformer.StatefulTransformer,artan.filter.filter_params.KalmanFilterParams,artan.filter.filter_params.HasMultipleModelAdaptiveEstimationEnabled,artan.filter.filter_params.HasMultipleModelMeasurementWindowDuration,artan.utils.ArtanJavaMLReadable,pyspark.ml.util.JavaMLWritableLinear Kalman Filter, implemented with a stateful spark Transformer for running parallel filters /w spark dataframes. Transforms an input dataframe of noisy measurements to dataframe of state estimates using stateful spark transformations, which can be used in both streaming and batch applications.
Assuming a state vector
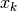 with size stateSize, and measurements vector
with size stateSize, and measurements vector 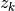 with size measurementSize, below parameters can be specified.
with size measurementSize, below parameters can be specified.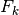 , process model, matrix with dimensions stateSize x stateSize
, process model, matrix with dimensions stateSize x stateSize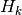 , measurement model, matrix with dimensions stateSize x measurementSize
, measurement model, matrix with dimensions stateSize x measurementSize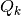 , process noise covariance, matrix with dimensions stateSize x stateSize
, process noise covariance, matrix with dimensions stateSize x stateSize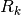 , measurement noise covariance, matrix with dimensions measurementSize x measurementSize
, measurement noise covariance, matrix with dimensions measurementSize x measurementSize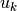 , optional control vector, vector with size controlSize
, optional control vector, vector with size controlSize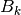 , optional control model, matrix with dimensions stateSize x controlSize
, optional control model, matrix with dimensions stateSize x controlSize
Linear Kalman Filter will predict & estimate the state according to following state and measurement equations.
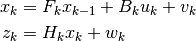
Where
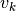 and
and 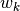 are noise vectors drawn from zero mean,
are noise vectors drawn from zero mean,
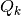 and
and 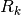 covariance distributions.
covariance distributions.The default values of system matrices will not give you a functioning filter, but they will be initialized with reasonable values given the state and measurement sizes. All of the inputs to the filter can be specified with a dataframe column which will allow you to have different value across measurements/filters, or you can specify a constant value across all measurements/filters.
artan.filter.recursive_least_squares_filter module¶
-
class
artan.filter.recursive_least_squares_filter.RecursiveLeastSquaresFilter[source]¶ Bases:
artan.state.stateful_transformer.StatefulTransformer,artan.filter.filter_params.HasInitialStateMean,artan.filter.filter_params.HasInitialStateMeanCol,artan.filter.recursive_least_squares_filter._HasForgettingFactor,artan.filter.recursive_least_squares_filter._HasRegularizationMatrix,artan.filter.recursive_least_squares_filter._HasRegularizationMatrixCol,pyspark.ml.param.shared.HasLabelCol,pyspark.ml.param.shared.HasFeaturesCol,artan.filter.filter_params.HasInitialStateDistributionCol,artan.filter.filter_params.HasStateSize,artan.utils.ArtanJavaMLReadable,pyspark.ml.util.JavaMLWritableRecursive formulation of least squares with exponential weighting & regularization, implemented with a stateful spark Transformer for running parallel filters /w spark dataframes. Transforms an input dataframe of observations to a dataframe of model parameters using stateful spark transformations, which can be used in both streaming and batch applications.
Let
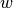 denote the model parameters and
denote the model parameters and 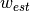 denote our prior belief. RLS minimizes following
regularization and weighted SSE terms;
denote our prior belief. RLS minimizes following
regularization and weighted SSE terms;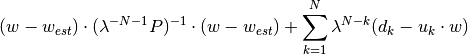
Where:
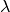 : forgetting factor, or exponential weighting factor. Between 0 and 1.
: forgetting factor, or exponential weighting factor. Between 0 and 1.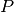 : regularization matrix. Smaller values increseas the weight of regularization term, whereas larger
: regularization matrix. Smaller values increseas the weight of regularization term, whereas larger- values increase the weight of weighted SSE term.
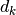 ,
, 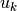 : label and features vector at time step k.
: label and features vector at time step k.
-
setFeatureSize(value)[source]¶ Set size of the features vector.
Parameters: value – Integer Returns: RecursiveLeastSquaresFilter
-
setFeaturesCol(value)[source]¶ Set features column. Default is “features”
Parameters: value – String Returns: RecursiveLeastSquaresFilter
-
setForgettingFactor(value)[source]¶ Set forgetting factor, exponentially weights the measurements based on its sequence.
Default value of 1.0 weights all measurements equally. With smaller values, recent measurements will have more weight. Generally set around 0.95 ~ 0.99
Parameters: value – Float >= 1.0 Returns: RecursiveLeastSquaresFilter
-
setInitialEstimate(value)[source]¶ Set initial estimate for model parameters. Default is zero vector.
Note that if this parameter is set through here, it will result in same initial estimate for all filters. For different initial estimates across filters, set the dataframe column for corresponding to initial estimate with setInitialEstimateCol.
Parameters: value – pyspark.ml.linalg.Vector with size (featuresSize) Returns: RecursiveLeastSquaresFilter
-
setInitialEstimateCol(value)[source]¶ Sets the column corresponding to initial estimates :param value: String :return: RecursiveLeastSquaresFilter
-
setInitialStateDistributionCol(value)[source]¶ Set the column corresponding to initial state distribution struct. The struct should have ‘mean’ field with a VectorType and ‘covariance’ field with a MatrixType. Overrides setInitialEstimateCol and setRegularizationMatrixCol parameters.
Parameters: value – String Returns: RecursiveLeastSquaresFilter
-
setLabelCol(value)[source]¶ Set label column. Default is “label”
Parameters: value – String Returns: RecursiveLeastSquaresFilter
-
setRegularizationMatrix(value)[source]¶ Set regularization matrix governing the influence of the initial estimate (prior). Larger values will remove regularization effect, making the filter behave like OLS.
Default is 10E5 * I
Note that if this parameter is set through here, it will result in same regularization for all filters. For different regularizations across filters, set the dataframe column for corresponding to regularization with setRegularizationMatrixCol.
Parameters: value – pyspark.ml.linalg.Matrix with size (featuresSize, featuresSize) Returns: RecursiveLeastSquaresFilter